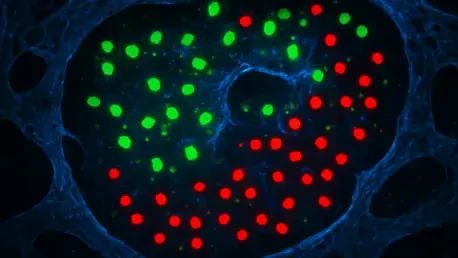
Scientists have successfully mapped the internal genetic blueprints of millions of individual cells, yet the precise physical coordinates of these cells within the living architecture of an organ often vanish the moment they are extracted for study. This “spatial gap” has long hindered the ability to link specific cellular behaviors to the precise microenvironments of complex organs like the brain or the kidney. To bridge this divide, researchers at the University Hospital Bonn and the University of Bonn developed the MERLIN algorithm, a sophisticated artificial intelligence tool designed to reconstruct the spatial origins of immune cells. By decoding the subtle “molecular memory” left by a cell’s environment, this breakthrough allows for a retrospective look at tissue geography, transforming how scientists interpret existing datasets and investigate localized disease processes.
Decoding the MERLIN Algorithm and Immune Cell Localization
The MERLIN algorithm emerged as a breakthrough AI tool designed specifically to address the loss of geographical data during standard laboratory procedures. In traditional research, isolating cells for analysis involves a process called dissociation, which effectively breaks down the organ into a cellular suspension. While this allows for deep genetic probing, it destroys the critical information regarding where those cells were originally positioned. MERLIN, which stands for Machine Learning-based Reconstruction of Localization in Immune cells, functions as a digital compass, allowing researchers to navigate back to the original tissue site by analyzing the “memory” encoded within the cell’s gene expression.
This technology focuses heavily on macrophages, the essential immune cells that reside within tissues to provide defense and maintain organ health. Because these cells are highly sensitive to their surroundings, they act as natural sensors of their environment. By utilizing advanced machine learning, the researchers demonstrated that it is possible to “read” the environmental history of these cells. This capability effectively closes the spatial gap in single-cell sequencing, providing a way to reconstruct the intricate physical maps of organs without needing to preserve the tissue structure during the initial sequencing phase.
The Significance of Spatial Context in Immunological Research
The rise of single-cell RNA sequencing (scRNA-seq) has provided a revolution in biological resolution, yet its major limitation remains its lack of a geographic map. When an organ is processed for scRNA-seq, the resulting data provides a list of cell types and their activity levels but offers no clue as to whether a specific cell was located in a nutrient-rich area or a zone of high stress. For structured tissues like the kidney or the brain, this lack of context is particularly detrimental. These organs are divided into highly specialized zones where oxygen levels, salt concentrations, and pressure vary significantly, and immune cells must adapt their behavior accordingly to survive and function.
The concept of “molecular memory” is central to understanding why spatial orientation is so vital for medical research. Environmental pressures, such as the extreme salt levels found in certain parts of the kidney or the unique chemical signaling in different brain regions, leave permanent transcriptomic signatures on the resident immune cells. These signatures act as a form of biological imprinting. When researchers can identify these signatures, they gain a deeper understanding of how the environment drives disease. For instance, a macrophage might behave differently in a high-oxygen zone compared to a low-oxygen zone, and knowing this distinction is key to developing treatments that target localized inflammation.
Research Methodology, Findings, and Implications
Methodology: Identifying Transcriptomic Echoes
The development of MERLIN relied on the identification of “transcriptomic echoes,” which are specific patterns of gene expression influenced by local tissue conditions. To ensure the algorithm could be used universally, the research team employed a rigorous training process involving diverse and independent datasets. This variety was essential to prevent computational overfitting, a common problem where an AI becomes too specialized to one specific set of data and fails to perform accurately in real-world scenarios. By using multiple sources, the algorithm learned to recognize consistent biological markers that signify specific locations within an organ.
The project also represented a significant integration of interdisciplinary techniques, blending expertise from the fields of immunology, nephrology, and bioinformatics. This collaboration allowed the team to filter genuine biological signals from the immense amount of genetic noise generated during sequencing. By focusing on the unique ways that immune cells respond to their environment, the researchers were able to create a set of “spatial fingerprints.” These fingerprints were then used to train the machine learning model to recognize where a cell came from based solely on its RNA profile, effectively turning gene expression data into a coordinate system.
Findings: Validating the Cellular Map
The research yielded impressive results, showing that MERLIN could successfully predict the localization of macrophages in both mouse models and human kidney samples. One of the most notable findings was the identification of specific gene markers that correspond to the renal medulla, an area of the kidney characterized by a high-salt environment. The algorithm was able to look at a cell’s genetic profile and accurately place it within this specific zone, confirming that the “salt stress” signatures were reliable indicators of location. This success demonstrated that the algorithm’s logic held true across different species, bridging the gap between basic research and clinical application.
To further validate the versatility of the algorithm, the team expanded their focus to the brain’s complex architecture. They applied MERLIN to map the positions of microglia, which are specialized immune cells of the central nervous system. Despite the brain’s vastly different chemical and physical makeup compared to the kidney, the algorithm remained highly accurate in reconstructing the spatial distribution of these cells. This suggested that the principle of environmental imprinting is a widespread biological phenomenon, making MERLIN a potentially universal tool for tissue mapping across various organ systems.
Implications: Advancing Precision Medicine
The implications of the MERLIN algorithm are vast, particularly regarding the ability to “re-mine” existing, publicly available datasets. Researchers around the world have already generated massive amounts of single-cell data that lack spatial context; MERLIN allows them to revisit this information and uncover hidden spatial insights without the need for new, expensive experiments. This capability could significantly accelerate the pace of discovery in many fields of medicine. By detailing how immune cells respond to localized stresses, scientists can gain a much clearer picture of how complex conditions like sepsis, diabetic nephropathy, and organ transplant injuries develop within specific tissue zones.
Furthermore, this technology marks a significant step forward for the advancement of precision medicine. By understanding how localized drug effects vary across different regions of an organ, clinicians can better predict how a patient might respond to a particular therapy. For example, a drug might be highly effective in one part of the kidney but cause adverse reactions in another. MERLIN provides the high-resolution data needed to observe these regional variations at a cellular level. This deeper understanding of human disease geography allows for the identification of new, highly targeted therapeutic targets that were previously invisible in non-spatial data.
Reflection and Future Directions
Reflection: Breaking Scientific Silos
The creation of MERLIN was the result of a concerted collaborative effort between several international institutions, highlighting the necessity of breaking scientific silos to solve the most complex problems in biology. By bringing together mathematicians, clinicians, and biologists, the project successfully addressed the long-standing “dissociation” barrier of traditional sequencing. The reflection on this process showed that maintaining biological context does not always require preserving physical tissue; instead, it requires smarter ways to interpret the data that remains. This project proved that the information needed to reconstruct the body’s internal maps was present all along, hidden within the transcriptome.
Future Directions: Expanding the Framework
The research team identified several exciting pathways for the future of the MERLIN framework. One primary goal is the expansion of the algorithm to include a wider variety of immune cell types beyond macrophages and microglia, potentially encompassing T-cells and neutrophils. There is also significant potential for using MERLIN in real-time clinical diagnostics to monitor the progression of diseases in three dimensions, providing a more comprehensive view of patient health. Additionally, future investigations will look into how “location memory” changes over time, particularly as an individual ages or as a chronic illness progresses, which could reveal why certain tissues become more vulnerable to disease later in life.
The Future of Spatial Transcriptomics in Clinical Medicine
The development of the MERLIN algorithm provided a necessary high-definition lens for the scientific community, turning static data into a dynamic map of human biology. Placing cells back into their anatomical context allowed for a more profound understanding of how the immune system interacts with its surroundings. This work demonstrated that the transcriptomic signatures left by oxygen, salt, and other environmental factors served as a reliable record of a cell’s life and function. By utilizing these records, the research established a new standard for how single-cell data is analyzed and interpreted in the context of the whole organ.
The project successfully concluded that the “memory” of a cell’s location is a fundamental aspect of its identity, which must be considered in any comprehensive study of disease. Moving forward, the focus shifted toward integrating these spatial insights into the development of next-generation therapies and diagnostic tools. The ability to map the geography of disease at a cellular level opened the door for more sophisticated medical interventions that account for the unique conditions of different tissue zones. Ultimately, the integration of AI-driven spatial reconstruction will continue to reshape the landscape of clinical medicine, leading to more personalized and effective treatments for a wide range of complex disorders.