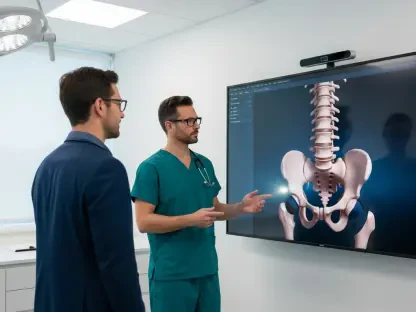
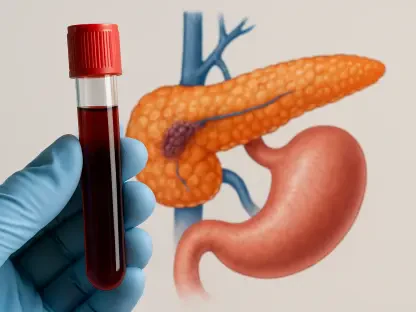
The medical community has long relied on visible motor symptoms like tremors and physical rigidity to diagnose Parkinson’s disease, yet these outward signs represent only the tip of a much deeper and more diverse biological struggle. Recent breakthroughs from the VIB-KU Leuven Center for Neuroscience have fundamentally challenged the traditional clinical consensus by revealing that what was once considered a single condition is actually a spectrum of distinct molecular disorders. By moving beyond simple observation, researchers have demonstrated that patients who appear to have the same illness may actually be suffering from entirely different cellular malfunctions. This paradigm shift is essential for understanding why treatments that work for one individual often fail for another, as the underlying genetic drivers vary significantly across the patient population. As we move through 2026, the focus has shifted from managing symptoms to decoding the specific biological signatures that define each person’s unique version of the disease.
Biological Diversity: The Failure of Universal Treatments
The historical approach to neurodegenerative care has frequently hit a wall because it ignores the striking genetic complexity inherent in the human population. While millions of people exhibit the classic signs of bradykinesia and gait instability, the internal mechanisms triggering these issues are rarely identical. Numerous different genes have been linked to the onset of the disease, with each mutation affecting a specific cellular pathway, from mitochondrial health to protein degradation. This diversity explains the recurring failure of “silver bullet” therapies that attempt to treat every patient with the same chemical compound. A medication designed to repair a specific metabolic pathway might be revolutionary for a small group of patients, yet it could prove completely ineffective for those whose symptoms are driven by a different genetic mechanism. This study highlights that the diversity of biological causes is too great for any single therapeutic strategy to succeed across the board.
Recognizing that Parkinson’s is a collection of related but unique disorders allows scientists to move toward a more sophisticated model of pathology. For too long, the medical field has grouped individuals based on their shared tremors rather than their shared biology, leading to inconsistent results in clinical trials and frustrated expectations for patients. The research emphasizes that the clinical appearance of a disease does not always correlate with its molecular origin, meaning that two people with identical tremors might require two completely different medications. By acknowledging this heterogeneity, the scientific community is finally addressing the “one-size-fits-all” failure that has hindered progress for decades. This shift in perspective is the necessary foundation for the next era of neurology, where the primary goal is to identify the root cause of an individual’s condition rather than just suppressing the physical symptoms that eventually emerge from those internal errors.
Advanced Methodology: Harnessing Genetics and Machine Learning
To investigate the hidden structures of the disease, the research team utilized Drosophila melanogaster models, which were genetically engineered to carry mutations across two dozen different genes associated with the condition. These biological models provided a controlled environment where the researchers could monitor the progression of the disease longitudinally, capturing an immense dataset of behavioral and physical data points. Unlike human studies, which are often clouded by lifestyle factors and varied environments, these models allowed for a precise analysis of how specific genetic changes influence biological health over time. By tracking the flies from the earliest stages of the disease, the team was able to identify subtle patterns in movement and health that would be impossible to detect in a standard clinical setting. This methodology provided the raw data necessary to begin categorizing the various forms of the disease based on their actual impact.
The massive amount of information gathered from these models was processed using unsupervised machine learning algorithms, a technique that allows for the detection of patterns without human interference or preconceived hypotheses. This unbiased approach is critical because it prevents the researchers from accidentally imposing their own clinical biases on the data, allowing the computer to find natural groupings based on biological reality. The algorithm analyzed how different genetic mutations affected the models’ behavior and health, eventually grouping them into clusters that shared similar molecular pathways. This integration of experimental biology and computational power represents a significant technological leap, as it enables the classification of complex diseases through hard data rather than subjective observation. This process has transformed the way researchers look at genetic mutations, proving that even disparate genes can belong to the same functional group.
Molecular Subdivisions: Mapping the Internal Landscape
The computational analysis led to a transformative discovery: the various genetic forms of Parkinsonism naturally cluster into two primary categories, which are further divided into five distinct molecular subdivisions. This represents the first time a comprehensive molecular classification of the disease has been achieved through behavioral outputs in a genetic model. It provides a long-awaited “map” of the internal landscape, proving that while different mutations eventually lead to the same clinical symptoms, they travel along very different biological routes to reach that destination. By identifying these five subdivisions, the research team has created a framework that allows for the categorization of the disease with a level of precision that was previously impossible. This clarity is essential for researchers who are attempting to develop therapies that target the specific cellular dysfunction driving the illness in a particular patient group.
Understanding these five subdivisions allows the medical community to move away from the vague umbrella of a single diagnosis and toward a more granular understanding of neurodegeneration. Each molecular subtype represents a unique set of challenges and biological failures, requiring a specialized approach to intervention. For example, a subtype characterized by mitochondrial stress will likely respond to different interventions than one defined by protein folding issues. By defining these signatures, the research provides a prerequisite for precision medicine, ensuring that clinicians can eventually match the right treatment to the right biological profile. This discovery effectively ends the era of treating the disease as a monolith and starts an era where the molecular “signature” of a patient is the most important factor in their care. The focus is no longer on the tremor itself, but on the specific biological error that created it.
Precision Medicine: Future Insights and Practical Solutions
To demonstrate the practical utility of these findings, the research team conducted a series of pharmacological tests on the different fly subtypes they had identified. They discovered that specific drug compounds could successfully reverse symptoms in one molecular subgroup while having absolutely no effect on others. This finding provides a definitive explanation for the high failure rate of many Parkinson’s drugs in human clinical trials. If a trial includes a diverse mix of patients from all five molecular subgroups, the positive results in one group are often statistically overwhelmed by the lack of response in the other four. This suggests a future trend where clinical research must stratify patients by molecular subtype before a trial begins. By ensuring that drugs are tested only on the populations they are biologically designed to help, the pharmaceutical industry can significantly improve the success rate of new neurodegenerative treatments.
Beyond the immediate impact on Parkinson’s care, this data-driven framework established a template for studying other complex conditions like Alzheimer’s or ALS. The transition toward precision medicine promised better patient outcomes by allowing for personalized diagnostic tests and targeted therapies. In the time following this research, the focus moved to identifying human biomarkers that correspond to these five specific subtypes, which would allow doctors to use simple tests to prescribe the most effective medications. The shift toward molecular classification reduced the incidence of side effects from ineffective treatments and improved the general quality of life for millions. By integrating advanced computational techniques with experimental biology, scientists opened a new door in the quest to solve complex puzzles in human health. This approach prioritized biological truth over clinical appearance, laying the groundwork for a more efficient and compassionate medical system.