I’m thrilled to sit down with Ivan Kairatov, a renowned biopharma expert with extensive experience in research and development, and a deep understanding of technological innovation in the medical field. Today, we’re diving into a groundbreaking advancement in sepsis detection that promises to transform patient care by drastically reducing diagnosis time. In our conversation, Ivan shares insights on how this new method works, why speed is so vital in treating sepsis, the specific challenges and successes in detecting various bacteria, and the broader implications for hospital practices and antibiotic use. Let’s explore how this innovation could save countless lives.
Can you start by giving us an overview of this exciting new method for detecting sepsis infections, and what sets it apart from traditional approaches?
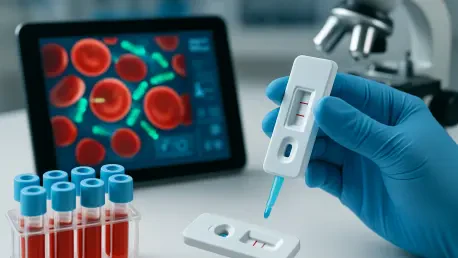
Absolutely, I’m glad to share details on this. This new method is a revolutionary approach to diagnosing sepsis by identifying bloodstream infections in just two hours, compared to the traditional bacteria culturing process that often takes at least a day or more in hospital labs. It uses a combination of centrifugation to separate bacteria from blood cells, automated microscopy for detection, and artificial intelligence software to analyze the results. What makes it stand out is its ability to bypass the long incubation periods of conventional methods, providing actionable insights almost immediately.
Why is getting a diagnosis so quickly such a critical factor when it comes to sepsis?
Speed is everything with sepsis because it’s truly a race against time. With every hour of delayed treatment in cases of septic shock, a patient’s survival rate drops by about 8 percent. That’s a staggering statistic. A faster diagnosis means we can start the right treatment sooner, preventing the rapid deterioration that can occur. Cutting the wait from a day to just two hours can literally be the difference between life and death for many patients.
Could you walk us through the nuts and bolts of how this diagnostic process actually functions?
Of course. The process begins with a technique we call ‘smart centrifugation,’ where a blood sample is spun with a special agent that causes bacteria to float upwards while blood cells sink. This creates a clear liquid layer containing just the bacteria. That liquid is then injected into a chip with tiny microscale channels and traps that capture the bacteria. From there, automated time-lapse microscopy takes images of any bacterial growth, and AI-driven software analyzes these images to identify the pathogens. It’s a highly precise and streamlined process that eliminates much of the manual waiting and guesswork.
What types of bacteria has this method been able to detect so far, and how effective has it been in testing?
In testing, the method has shown great success with several key bacteria associated with sepsis, including E. coli, K. pneumoniae, and E. faecalis, detecting them at very low, clinically relevant levels—between 7 to 32 colony-forming units per milliliter of blood. These results are incredibly promising. However, it struggled with Staphylococcus aureus, which tends to hide in blood clots, making it harder to isolate. We’re actively working on refining the technique to address this challenge and ensure broader applicability.
How does this faster detection method impact the way antibiotics are prescribed for sepsis patients?
It’s a game-changer in terms of antibiotic stewardship. Currently, when sepsis is suspected, doctors often start patients on broad-spectrum antibiotics as a precaution, which can be toxic, disrupt beneficial gut bacteria, and contribute to antibiotic resistance. With this method, we can identify the specific pathogen in hours, not days, allowing doctors to switch to a targeted antibiotic much sooner. This reduces risks to the patient and helps combat the growing problem of resistant strains.
What kind of transformation do you envision this technology bringing to hospital labs and overall patient care?
I believe this could fundamentally change how hospital labs operate. Instead of waiting two to four days to confirm the right treatment, we’re talking about potentially reducing that to just four to six hours. This not only improves patient outcomes but also eases the burden on lab resources. As for becoming a standard tool, I think with further validation and scaling, it has the potential to be widely adopted in hospitals within the next few years, especially as the technology becomes more accessible.
Can you share some insights into the collaborative efforts that made this project possible?
This breakthrough wouldn’t have happened without strong collaboration between multidisciplinary teams. Experts in microfluidics, biomedical systems, and digital imaging came together, combining their unique skills to tackle different aspects of the challenge. It was a synergy of technical expertise in separating and detecting bacteria, paired with advanced software development for analysis. This kind of teamwork is essential for translating complex research into real-world solutions.
Looking ahead, what is your forecast for the future of rapid diagnostic technologies like this in the fight against sepsis?
I’m incredibly optimistic. I think we’re on the cusp of a new era where rapid diagnostics will become the norm, not the exception, in managing critical conditions like sepsis. As we refine these technologies—improving their accuracy across more pathogens and making them more cost-effective—I foresee them being integrated into routine clinical workflows worldwide. This could lead to a significant reduction in sepsis mortality rates and set a precedent for how we approach other time-sensitive medical challenges. The potential to save lives on a global scale is truly inspiring.